Reject Option based Classifier is post-processing bias mitigation method. Method changes labels of favorable, privileged and close to cutoff observations to unfavorable and the opposite for unprivileged observations (changing unfavorable and close to cutoff observations to favorable, more in details). By this potentially wrongfully labeled observations are assigned different labels. Note that in y in DALEX explainer 1 should indicate favorable outcome.
roc_pivot(explainer, protected, privileged, cutoff = 0.5, theta = 0.1)Arguments
| explainer | created with |
|---|---|
| protected | factor, protected variables with subgroups as levels (sensitive attributes) |
| privileged | factor/character, level in protected denoting privileged subgroup |
| cutoff | numeric, threshold for all subgroups |
| theta | numeric, variable specifies maximal euclidean distance to cutoff resulting ing label switch |
Value
DALEX explainer with changed y_hat. This explainer should be used ONLY by fairmodels as it contains unchanged
predict function (changed predictions (y_hat) can possibly be invisible by DALEX functions and methods).
Details
Method implemented implemented based on article (Kamiran, Karim, Zhang 2012). In original implementation labels should be switched. Due to specific DALEX methods
probabilities (y_hat) are assigned value in equal distance but other side of cutoff. The method changes explainers y_hat values in two cases.
1. When unprivileged subgroup is within (cutoff - theta, cutoff)
2. When privileged subgroup is within (cutoff, cutoff + theta)
References
Kamiran, Karim, Zhang 2012 https://ieeexplore.ieee.org/document/6413831/ ROC method
Examples
data("german")
data <- german
data$Age <- as.factor(ifelse(data$Age <= 25, "young", "old"))
y_numeric <- as.numeric(data$Risk) - 1
lr_model <- stats::glm(Risk ~ ., data = data, family = binomial())
lr_explainer <- DALEX::explain(lr_model, data = data[, -1], y = y_numeric)
#> Preparation of a new explainer is initiated
#> -> model label : lm ( default )
#> -> data : 1000 rows 9 cols
#> -> target variable : 1000 values
#> -> predict function : yhat.glm will be used ( default )
#> -> predicted values : No value for predict function target column. ( default )
#> -> model_info : package stats , ver. 4.1.1 , task classification ( default )
#> -> predicted values : numerical, min = 0.1188633 , mean = 0.7 , max = 0.9782676
#> -> residual function : difference between y and yhat ( default )
#> -> residuals : numerical, min = -0.9631188 , mean = 3.356085e-16 , max = 0.8419051
#> A new explainer has been created!
fobject <- fairness_check(lr_explainer,
protected = data$Age,
privileged = "old"
)
#> Creating fairness classification object
#> -> Privileged subgroup : character ( Ok )
#> -> Protected variable : factor ( Ok )
#> -> Cutoff values for explainers : 0.5 ( for all subgroups )
#> -> Fairness objects : 0 objects
#> -> Checking explainers : 1 in total ( compatible )
#> -> Metric calculation : 13/13 metrics calculated for all models
#> Fairness object created succesfully
plot(fobject)
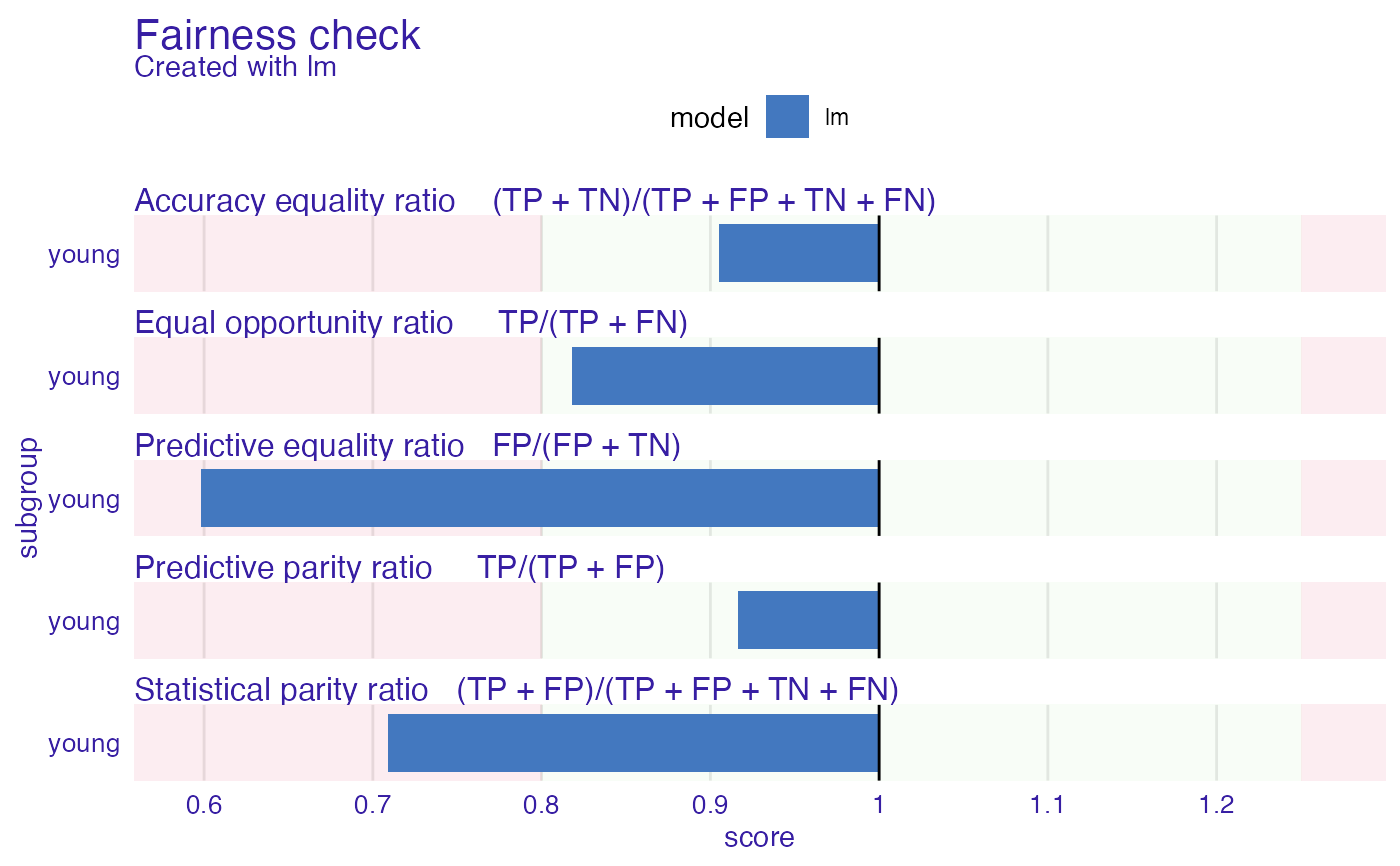 lr_explainer_fixed <- roc_pivot(lr_explainer,
protected = data$Age,
privileged = "old"
)
fobject2 <- fairness_check(lr_explainer_fixed, fobject,
protected = data$Age,
privileged = "old",
label = "lr_fixed"
)
#> Creating fairness classification object
#> -> Privileged subgroup : character ( Ok )
#> -> Protected variable : factor ( Ok )
#> -> Cutoff values for explainers : 0.5 ( for all subgroups )
#> -> Fairness objects : 1 object ( compatible )
#> -> Checking explainers : 2 in total ( compatible )
#> -> Metric calculation : 13/13 metrics calculated for all models
#> Fairness object created succesfully
fobject2
#>
#> Fairness check for models: lr_fixed, lm
#>
#> lr_fixed passes 4/5 metrics
#> Total loss : 0.7380378
#>
#> lm passes 3/5 metrics
#> Total loss : 1.051602
#>
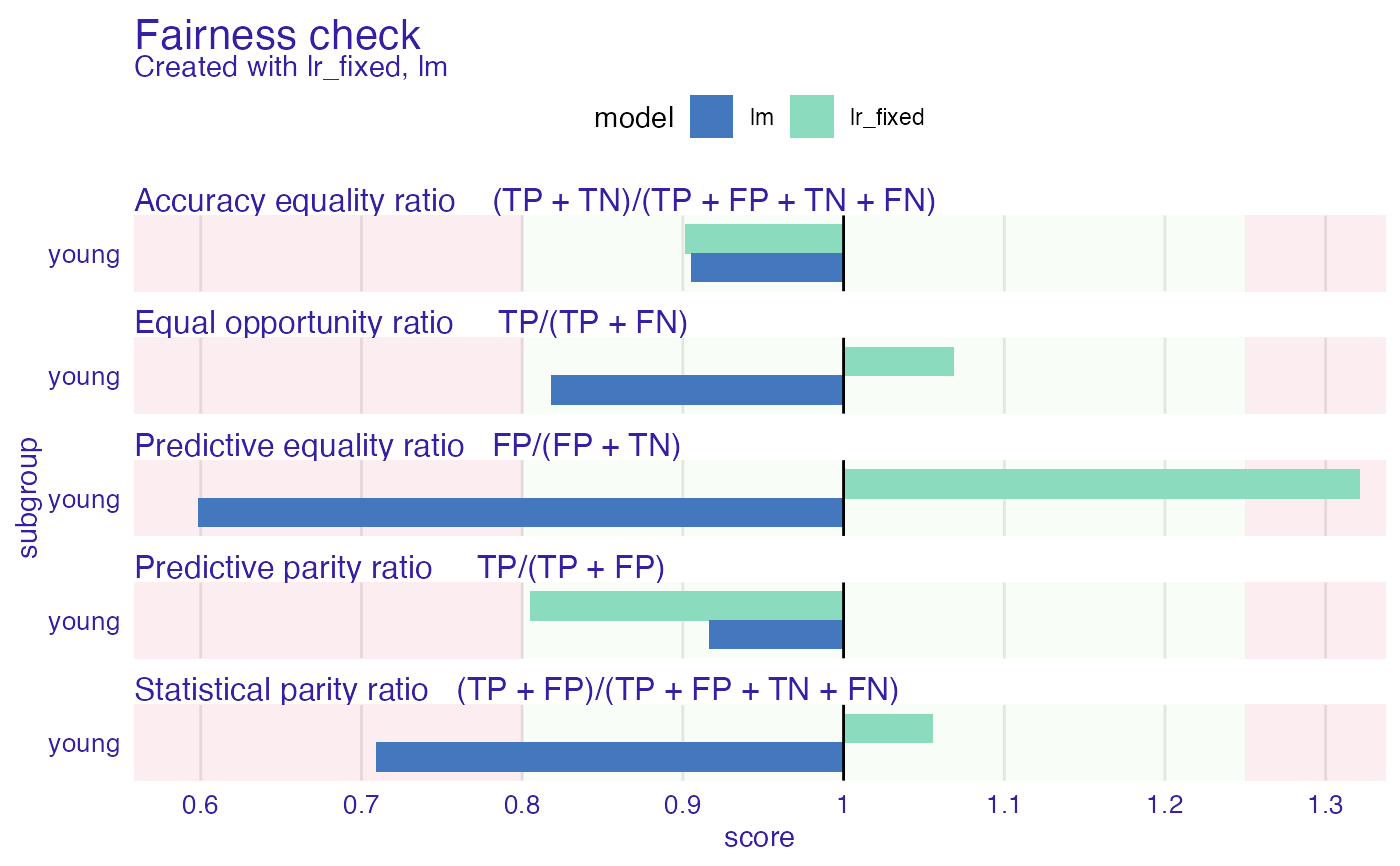
plot(fobject2)
lr_explainer_fixed <- roc_pivot(lr_explainer,
protected = data$Age,
privileged = "old"
)
fobject2 <- fairness_check(lr_explainer_fixed, fobject,
protected = data$Age,
privileged = "old",
label = "lr_fixed"
)
#> Creating fairness classification object
#> -> Privileged subgroup : character ( Ok )
#> -> Protected variable : factor ( Ok )
#> -> Cutoff values for explainers : 0.5 ( for all subgroups )
#> -> Fairness objects : 1 object ( compatible )
#> -> Checking explainers : 2 in total ( compatible )
#> -> Metric calculation : 13/13 metrics calculated for all models
#> Fairness object created succesfully
fobject2
#>
#> Fairness check for models: lr_fixed, lm
#>
#> lr_fixed passes 4/5 metrics
#> Total loss : 0.7380378
#>
#> lm passes 3/5 metrics
#> Total loss : 1.051602
#>
plot(fobject2)