Ceteris paribus cutoff is way to check how will parity loss behave if we changed only cutoff in one subgroup. It plots object of class ceteris_paribus_cutoff. It might have two types - default and cumulated. Cumulated sums metrics and plots it all in one plot. When default one is used all chosen metrics will be plotted for each model.
# S3 method for ceteris_paribus_cutoff
plot(x, ...)Arguments
| x | ceteris_paribus_cutoff object |
|---|---|
| ... | other plot parameters |
Value
ggplot2 object
Examples
data("compas")
# positive outcome - not being recidivist
two_yr_recidivism <- factor(compas$Two_yr_Recidivism, levels = c(1, 0))
y_numeric <- as.numeric(two_yr_recidivism) - 1
compas$Two_yr_Recidivism <- two_yr_recidivism
lm_model <- glm(Two_yr_Recidivism ~ .,
data = compas,
family = binomial(link = "logit")
)
explainer_lm <- DALEX::explain(lm_model, data = compas[, -1], y = y_numeric)
#> Preparation of a new explainer is initiated
#> -> model label : lm ( default )
#> -> data : 6172 rows 6 cols
#> -> target variable : 6172 values
#> -> predict function : yhat.glm will be used ( default )
#> -> predicted values : No value for predict function target column. ( default )
#> -> model_info : package stats , ver. 4.1.1 , task classification ( default )
#> -> predicted values : numerical, min = 0.004522979 , mean = 0.5448801 , max = 0.8855426
#> -> residual function : difference between y and yhat ( default )
#> -> residuals : numerical, min = -0.8822826 , mean = -5.07018e-13 , max = 0.9767658
#> A new explainer has been created!
fobject <- fairness_check(explainer_lm,
protected = compas$Ethnicity,
privileged = "Caucasian"
)
#> Creating fairness classification object
#> -> Privileged subgroup : character ( Ok )
#> -> Protected variable : factor ( Ok )
#> -> Cutoff values for explainers : 0.5 ( for all subgroups )
#> -> Fairness objects : 0 objects
#> -> Checking explainers : 1 in total ( compatible )
#> -> Metric calculation : 11/13 metrics calculated for all models ( 2 NA created )
#> Fairness object created succesfully
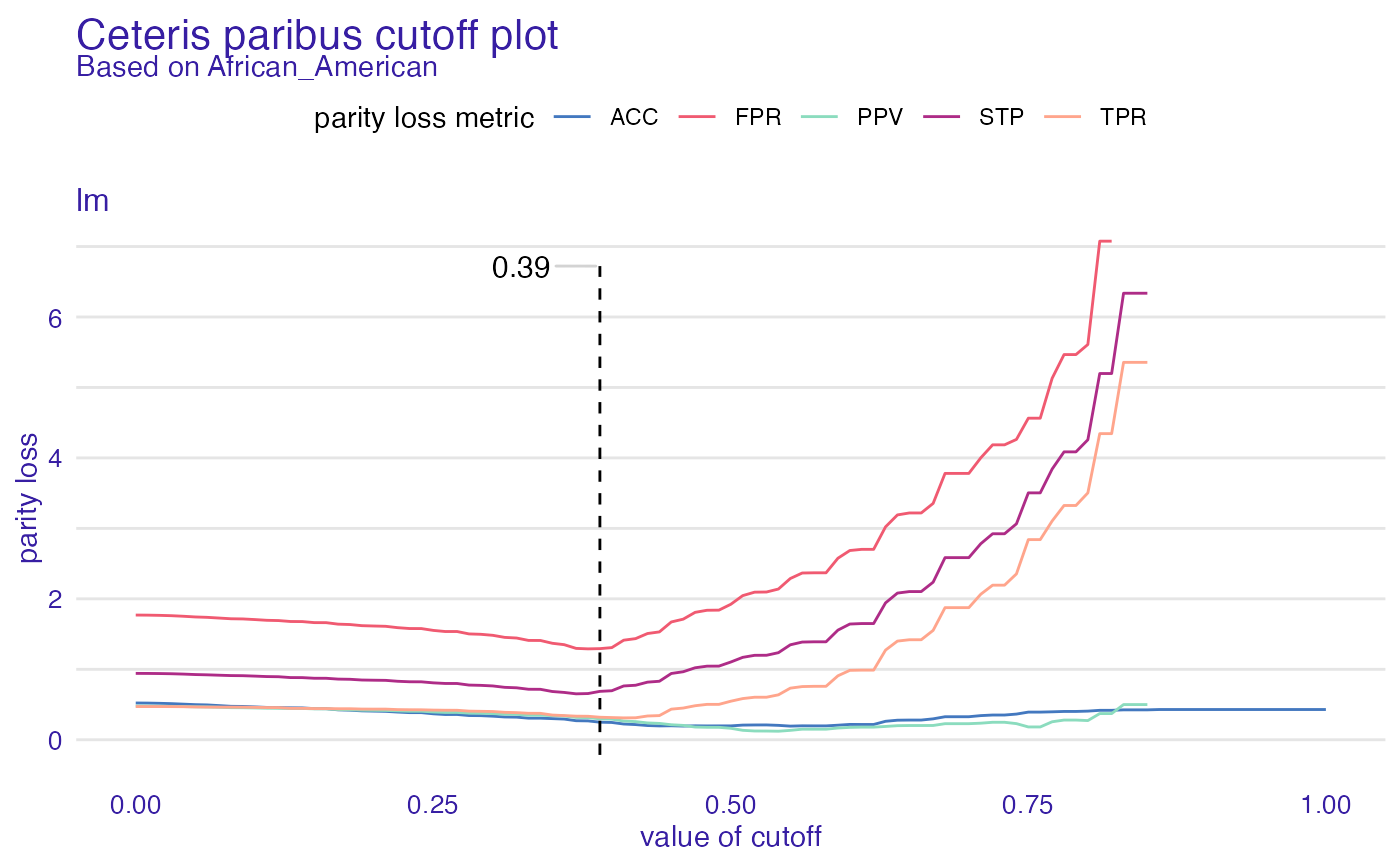
cpc <- ceteris_paribus_cutoff(fobject, "African_American")
plot(cpc)
#> Warning: Removed 63 row(s) containing missing values (geom_path).
 # \donttest{
rf_model <- ranger::ranger(Two_yr_Recidivism ~ .,
data = compas,
probability = TRUE,
num.trees = 200
)
explainer_rf <- DALEX::explain(rf_model, data = compas[, -1], y = y_numeric)
#> Preparation of a new explainer is initiated
#> -> model label : ranger ( default )
#> -> data : 6172 rows 6 cols
#> -> target variable : 6172 values
#> -> predict function : yhat.ranger will be used ( default )
#> -> predicted values : No value for predict function target column. ( default )
#> -> model_info : package ranger , ver. 0.13.1 , task classification ( default )
#> -> predicted values : numerical, min = 0.153945 , mean = 0.5452013 , max = 0.8650518
#> -> residual function : difference between y and yhat ( default )
#> -> residuals : numerical, min = -0.847902 , mean = -0.0003212319 , max = 0.7775046
#> A new explainer has been created!
fobject <- fairness_check(explainer_lm, explainer_rf,
protected = compas$Ethnicity,
privileged = "Caucasian"
)
#> Creating fairness classification object
#> -> Privileged subgroup : character ( Ok )
#> -> Protected variable : factor ( Ok )
#> -> Cutoff values for explainers : 0.5 ( for all subgroups )
#> -> Fairness objects : 0 objects
#> -> Checking explainers : 2 in total ( compatible )
#> -> Metric calculation : 8/13 metrics calculated for all models ( 5 NA created )
#> Fairness object created succesfully
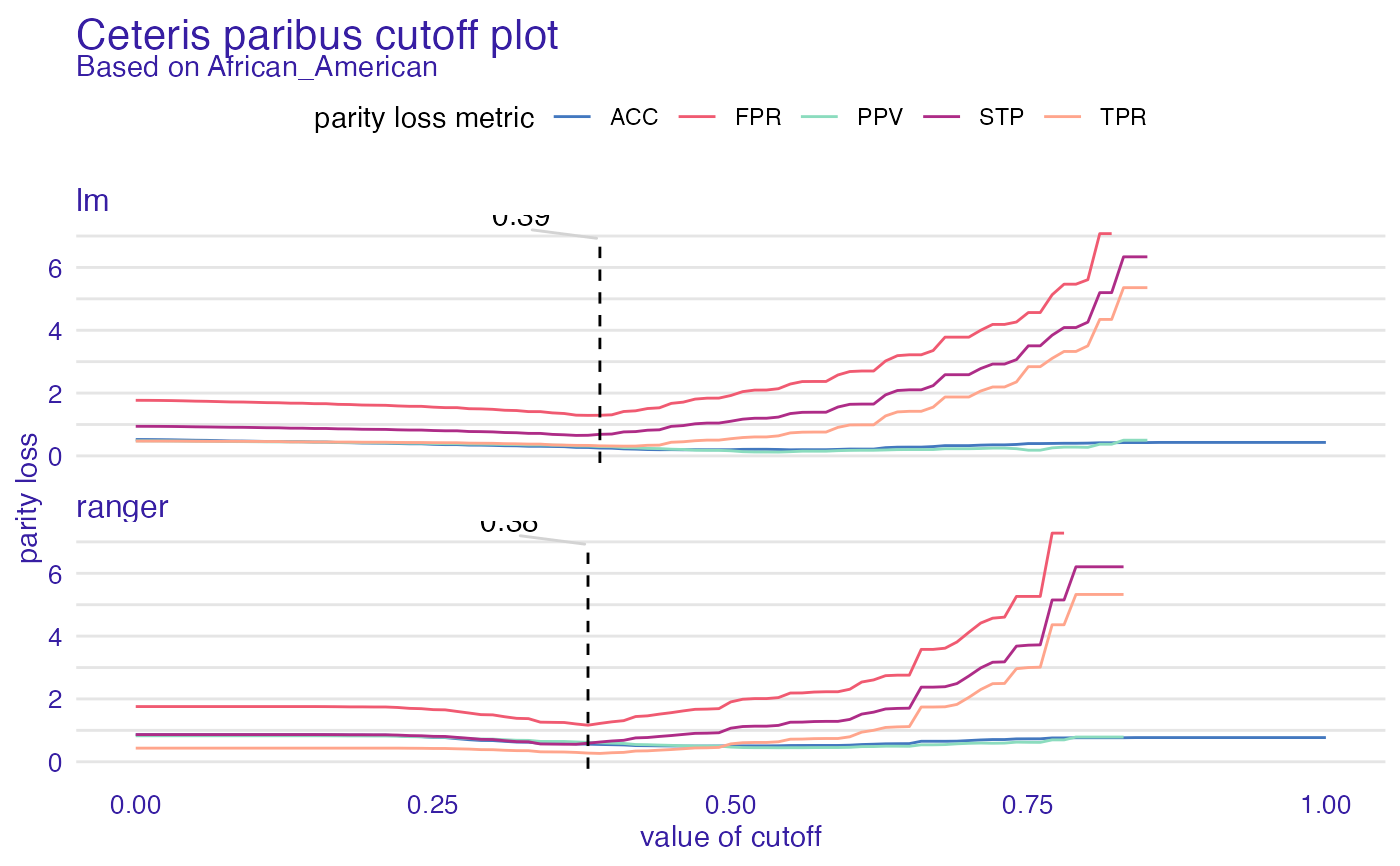
cpc <- ceteris_paribus_cutoff(fobject, "African_American")
plot(cpc)
#> Warning: Removed 73 row(s) containing missing values (geom_path).
# \donttest{
rf_model <- ranger::ranger(Two_yr_Recidivism ~ .,
data = compas,
probability = TRUE,
num.trees = 200
)
explainer_rf <- DALEX::explain(rf_model, data = compas[, -1], y = y_numeric)
#> Preparation of a new explainer is initiated
#> -> model label : ranger ( default )
#> -> data : 6172 rows 6 cols
#> -> target variable : 6172 values
#> -> predict function : yhat.ranger will be used ( default )
#> -> predicted values : No value for predict function target column. ( default )
#> -> model_info : package ranger , ver. 0.13.1 , task classification ( default )
#> -> predicted values : numerical, min = 0.153945 , mean = 0.5452013 , max = 0.8650518
#> -> residual function : difference between y and yhat ( default )
#> -> residuals : numerical, min = -0.847902 , mean = -0.0003212319 , max = 0.7775046
#> A new explainer has been created!
fobject <- fairness_check(explainer_lm, explainer_rf,
protected = compas$Ethnicity,
privileged = "Caucasian"
)
#> Creating fairness classification object
#> -> Privileged subgroup : character ( Ok )
#> -> Protected variable : factor ( Ok )
#> -> Cutoff values for explainers : 0.5 ( for all subgroups )
#> -> Fairness objects : 0 objects
#> -> Checking explainers : 2 in total ( compatible )
#> -> Metric calculation : 8/13 metrics calculated for all models ( 5 NA created )
#> Fairness object created succesfully
cpc <- ceteris_paribus_cutoff(fobject, "African_American")
plot(cpc)
#> Warning: Removed 73 row(s) containing missing values (geom_path).
 # }
# }